Department of Molecular Biology
Notice: The 12th 3R+3C International Symposium (Nov. 18-22, 2024)
Faculty members
Tsutomu Katayama, Ph.D. , Professor
Shogo Ozaki, Ph.D., Associate Professor
Kazutoshi Kasho, Ph. D., Assistant Professor
Related, brief history
(Hironori Kawakami, Ph.D., a previous Reseach Associate ---> an Uehara Foundation (later, a JSPS) post-doctoral fellow in Dr. Bruce Stillman Laboratory of CSHL ---> Assistant Professor from March 2012--->Associate Professor of Sanyo Onoda-City University
(Msayuki Su'etsugu, Ph.D., a previous Reseach Associate ---> a JSPS post-doctoral fellow in Dr. Jeff Errington Laboratory of Newcastle University (UK) ---> Assistant Professor from October 2011 ----> From April 2013, Associate Professor of Rikkyo University ----> Professor of Rikkyo University
(Kazuyuki Fujimitsu, Ph.D., a previous Reseach Associate ----> a post-doctoral fellow in Dr. Yamano Laboratory in UCL Cancer Institute, University College London .)
(Shogo Ozaki, Ph.D., a previous Reseach Associate ----> an Uehara Foundation post-doctoral fellow in Dr. Urs Jenal Laboratory in Biozentrum of University of Basel ----> Associate Professor (present)
(Kenji Keyamura, Ph.D., a previous Reseach Associate ----> an Assistant Professor of Gakushuin University in Tokyo
(Yasunori Noguchi, Ph.D. a previous Reseach Associate ----> a post-doctoral fellow in Dr. Christian Speck Laboratory in UCL (UK) ----> an Assistant Professor of Dr. Masatoshi Fujita Laboratory in Kyushu University
(Kazutoshi Kasho, Ph.D. ----> a JSPS post-doctoral fellow in Umea University (Sweden) ---->Assistant Professor (present)
Research
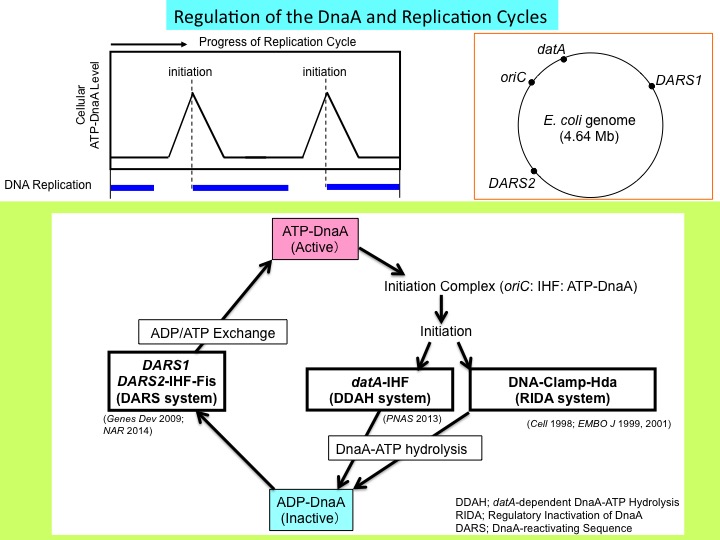
In the cell cycle progression, chromosomal DNA is replicated only once at a specific time by the carefully controlled molecular switch for replicational initiation. If this regulation is interfered with, various cell defects occur, such as abnormal chromosomes, inhibition of cell division, and growth of abnormal cells. Thus, a study on this regulatory mechanism is of significance as a basis for the developments of antibiotics and anticancer drugs. We have shown that a protein (DnaA) initiating E. coli chromosomal replication is inactivated by timely and direct interaction with a subunit (beta subunit) of the chromosomal replicase (DNA polymerase III holoenzyme). This interaction depends on loading the subunit onto DNA. This conformational change occurs for the nucleotide-polymerizing action of the replicase after the initiation reaction by DnaA. Thus, during the cell cycle, the initiation protein is inactivated after initiation of chromosomal replication in a DNA replication-coupled feedback manner. We have termed this regulatory system RIDA (Regulatory inactivation of DnaA). In addition, we have also shown that the chromosomal DNA element (datA) is activated in a specific time after replication initiation for inactivating DnaA. Reactivation of DnaA occurs before the next round of the replication cycle, as we have shown that the chromosomal DNA element (DARS2) is activated in a timely manner for re-activating DnaA. We are further investigating molecular mechanisms in this DnaA-activity cycle including timely inactivation and activation, in addition to molecular mechanisms in replication initiation sustained by higher-order complexes including DnaA and the chromosomal replication orign oriC. We have shown that localized DNA bending (looping) in oriC promotes DnaA-dependent DNA unwinding (ssDUE recruitment mechanism).
Important references
Text
Benjamin Lewin (2004) "GENES VIII, International Edition"
ü@CHAPTER 14 ü@DNA Replicationü@ü@(RIDA is described on p410-411)
Review
Kasho, K., Ozaki, S., and Katayama, T. (2023)
IHF and Fis as Escherichia coli cell cycle regulators: Activation of the replication origin oriC and the regulatory cycle of the DnaA initiator
Int. J. Mol. Sci. 24(14); 11572
https://doi.org/10.3390/ijms241411572Katayama, T. (2017)
Initiation of DNA replication at the chromosomal origin of E. coli, oriC
A chapter in the book entitled 'DNA Replicationn: From Old Principles to New Discovery' (Hisao Masai and Marco Foiani, Eds): Adv Exp Med Biol.1042, SpringerNature pp 79-98.
doi: 10.1007/978-981-10-6955-0_4
Katayama, T., Kasho, K., and Kawakami, H. (2017)
The DnaA cycle in Escherichia coli: activation, function, and inactivation of the initiator protein
Front. Microbiol. 8:2496. doi: 10.3389/fmicb.2017.02496Skarstad, K. and Katayama, T. (2013)
Regulating DNA Replication in Bacteria
In "DNA Replication", Edited by Bell, S.D., Me'chali, M, and DePamphilis, M.L., Cold Spring Harbor Laboratory PressKatayama, T., Ozaki, S., Keyamura, K. and Fujimitsu, K. (2010)
Regulation of the replication cycle: Conserved and diverse regulatory systems for DnaA and oriC
Nature Rev. Microbiol. 8(3):163-170Kawakami, H. and Katayama, T. (2010)
DnaA, ORC, and Cdc6: Similarity beyond the domains of life and diversity
Biochem. Cell Biol. 88(22): 49-62Ozaki, S., and Katayama, T. (2009)
DnaA structure, function, and dynamics in the initiation at the chromosomal origin
Plasmid 62: 71- 82Katayama, T.(2001)
Feedback controls restrain the initiation of Escherichia coli chromosomal replication.
Mol. Microbiol.41(1),9-18.
Selected original papersYoshida, R., Ozaki, S., Kawakami, H., and Katayama, T. (2023)
Single-stranded DNA recruitment mechanism in replication origin unwinding by DnaA initiator protein and HU, an evolutionary ubiquitous nucleoid protein
Nucleic Acids Res. 51(12): 6286-6306
https://doi.org/10.1093/nar/gkad389
Sakiyama, Y., Nagata, M., Yoshida, R. (equal contributors), Kasho, K., Ozaki, S., and Katayama. T. (2022)
Concerted actions of DnaA complexes with DNA unwinding sequences within and flanking replication origin oriC promote DnaB helicase loading
J. Biol. Chem. 298(6): 102051
doi: 10.1016/j.jbc.2022.102051
Ozaki, S., Wang, D., Wakasugi, Y., Itani, N., and Katayama, T. (2022)
The Caulobacter crescentus DciA promotes chromosome replication through topological loading of the DnaB replicative helicase at replication forks
Nucleic Acids Res. 50(22):12896-12912
doi.org/10.1093/nar/gkac1146Miyoshi, K., Tatsumoto, Y., Ozaki, S., and Katayama, T. (2021)
Negative feedback for DARS2-Fis complex by ATP-DnaA supports the cell cycle-coordinated regulation for chromosome replication
Nucleic Acids Res. 49(22): 12820-12835 Faculty Opinions Recomendation
doi: org/10.1093/nar/gkab1171
Taniguchi, S., Kasho, K., Ozaki, S., and Katayama, T. (2019)
Escherichia coli CrfC protein, a nucleoid partition factor, localizes to nucleoid poles via the activities of specific nucleoid-associated proteins.
Front. Microbiol. 10: 72. F1000Prime Recomendation
doi: 10.3389/fmicb.2019.00072
Sugiyama, R., Kasho, K. (equal contributors), Miyoshi, K., Ozaki, S., Kagawa, W., Kurumizaka, H., and Katayama, T. (2019)
A novel mode of DnaA-DnaA interaction promotes ADP dissociation for reactivation of replication initiation activity
Nucleic Acids Res. 47(21):11209-11224 F1000Prime Recomendation
doi: 10.1093/nar/gkz795
Shimizu, M., Noguchi, Y., Sakiyama, Y., Kawakami, H., Katayama, T.*, and Takada, S.* (*Co-corresponding authors) (2016)
Near-atomic structural model for bacterial DNA replication initiation complex and its functional insights
Proc. Natl. Acad. Sci. USA, 113(50); E8021-E8030 doi:10.1073/pnas.1609649113Kawakami, H., Ohashi, E., Kanamoto, S., Tsurimoto, T., and Katayama, T. (2015)
Specific binding of eukaryotic ORC to DNA replication origins depends on highly conserved basic residues
Sci. Rep. 5; 14929Ozaki, S., Matsuda, Y., Keyamura, K., Kawakami, H., Noguchi, Y., Kasho, K., Nagata, K., Masuda, T., Sakiyama, Y., and Katayama, T. (2013)
A replicase clamp-binding dynamin-like protein promotes colocalization of the nascent DNA strands and equipartitioning of chromosomes in E. coli
Cell Reportsüihttp://dx.doi.org/10.1016/j.celrep.2013.07.040Kasho K., and Katayama,T. (2013)
DnaA-binding locus datA promotes DnaA-ATP hydrolysis to enable cell cycle-coordinated replication initiation
Proc. Natl. Acad. Sci. USA 110, 946-941 F1000Prime RecomendationOzaki, S., and Katayama, T. (2012)
Highly organized DnaA-oriC complexes recruit the single-stranded DNA for replication initiation
Nucleic Acids Res. 40(4), 1648-1665Fujimitsu, K., Senriuchi, T., and Katayama, T. (2009)
Specific genomic sequences of E. coli promote replicational initiation by directly reactivating ADP-DnaA
Genes Dev. 23(10): 1221-1233 F1000Prime Recomendation (Perspectives, 23(10):1145-50)ü@Ozaki, S., Kawakami,H., Nakamura, K., Fujikawa, N., Kagawa, W., Park, S.-Y., Yokoyama, S., Kurumizaka, H., and Katayama, T. (2008)
A common mechanism for the ATP-DnaA-dependent formation of open complexes at the replication origin
J. Biol. Chem. 283(13), 8351-8362Keyamura, K., Fujikawa, N.(Equal contributors), Ishida, T., Ozaki, S., Su'etsugu, M., Fujimitsu, K., Kagawa, W., Yokoyama, S., Kurumizaka, H.* and Katayama, T.* (*Co-corresponding authos) (2007)
The interaction of DiaA and DnaA regulates the replication cycle in E. coli by directly promoting ATP-DnaA-specific initiation complexes
Genes Dev. 21, 2083-2099Abe, Y., Jo, T., Matsuda, Y., Matsunaga, C., Katayama, T.*, and Ueda, T.* (*Co-corresponding authors) (2007)
Structure and function of DnaA N-terminal domains: Specific sites and mechanisms in inter-DnaA interaction and in DnaB helicase loading on oriC
J. Biol. Chem. 282(24),17816-17827 JBC Paper of the Week
Kawakami, H., Keyamura, K., and Katayama, T. (2005)
Formation of an ATP-DnaA-specific initiation complex requires DnaA arginine-285, a conserved motif in the AAA+ protein family
J. Biol. Chem. 280(29), 27420-27430
Su'etsugu, M, Shimuta, T., Ishida, T., Kawakami, H. and Katayama, T. (2005)
Protein associations in DnaA-ATP hydrolysis mediated by the replicase clamp-Hda complex
J. Biol. Chem. 280(8), 6528-6536
Fujikawa, N., Kurumizaka, H., Nureki, O., Terada, T., Shirouzu, M., Katayama, T., and Yokoyama, S. (2003)
Structural basis of replication origin recognition by the DnaA protein
Nucleic Acid Res. 31(8), 2077-2086
Nishida, S., Fujimitsu, K., Sekimizu, K., Ohmura,T., Ueda, T., and Katayama, T.(2002)
A nucleotide switch in E.coli DnaA protein initiates chromosomal replication: Evidence from a mutant DnaA protein defective in regulatory ATP hydrolysis in vitro and in vivo.
J.Biol.Chem. 277(17),14986-14995
Kato,J. and Katayama, T.(2001)
Hda,a novel DnaA-related protein,regulates the replication cycle in Escherichia coli.
EMBO J. 20(15),4253-4262Kurokawa, K., Nishida, S., Emoto, A., Sekimizu, K.,and Katayama, T.(1999)
Replication cycle-coordinated change of the adenine nucleotide-bound forms of DnaA protein in Escherichia coli.
EMBO J. 18(23),6642-6652Katayama, T., Kubota, T., Kurokawa, K., Crooke, E.,and Sekimizu, K.(1998)
The initiation function of DnaAprotein is negatively regulated by the sliding clamp of the sliding clamp of the E.coli chromosomal replicase.
Cell 94(1),61-71